This notebook demonstrates how to use CstarSpecEngine to build a single domain configuration from domains.yml.
CstarSpecEngine Overview¶
CstarSpecEngine is the high-level interface for managing ROMS model configurations and builds. It provides methods to:
Generate single domains: Build one domain at a time using
generate_domain()Generate multiple domains: Build all domains from
domains.ymlusinggenerate_all()Run simulations: Execute model runs using
run_all()
The engine reads domain configurations from domains.yml and model specifications from models.yml, orchestrating the complete workflow from input generation through model compilation.
Domain Configuration¶
Domain configurations are defined in domains.yml. Each domain entry specifies:
Grid parameters: Resolution, size, location, vertical levels
Time range: Start and end dates for the simulation
Open boundaries: Which boundaries are open (north, south, east, west)
Partitioning: Parallel execution configuration
Model specification: Which model configuration to use (from
models.yml)
The domain name used here (_test-tiny) must match an entry in domains.yml.
Workflow Stages¶
The generate_domain() method executes the complete workflow:
PRECONFIG: Initialize blueprint and grid object
Source Data: Download and prepare required datasets (GLORYS, UNIFIED, SRTM15, etc.)
POSTCONFIG: Generate all input files (grid, initial conditions, forcing)
BUILD: Render configuration templates and compile the model executable
Pre-run: Prepare for execution (partitioning, etc.)
The method returns a CstarSpecBuilder instance that can be used to run the simulation.
Generate Domain¶
Create a CstarSpecEngine instance and generate a single domain. The domain name must match an entry in domains.yml.
Parameters:
domain_name: Name of the domain fromdomains.ymlclobber_inputs: IfTrue, overwrite existing input filesOther optional parameters:
clobber_source_data,partition_files,test,compile_time_settings,run_time_settings
%load_ext autoreload
%autoreload 2
from pathlib import Path
from glob import glob
import time
import cson_forge
import cstar.execution.handler as handler
import xarray as xrERROR 1: PROJ: proj_create_from_database: Open of /home/x-mlong/.local/share/mamba/envs/cson-forge-v0/share/proj failed
engine = cson_forge.CstarSpecEngine(domains_file="domains.yml")
builder = engine.generate_domain("ccs-12km", clobber_inputs=True)
builder✔️ Using existing GLORYS_REGIONAL file for 2024-01-01: cmems_mod_glo_phy_my_0.083deg_P1D-m_REGIONAL_ccs-12km_20240101.nc
✔️ Using existing GLORYS_REGIONAL file for 2024-01-02: cmems_mod_glo_phy_my_0.083deg_P1D-m_REGIONAL_ccs-12km_20240102.nc
✔️ TPXO dataset verified at: /anvil/projects/x-ees250129/cson-forge-data/source-data/TPXO/TPXO10.v2
✔️ Using existing BGC dataset: /anvil/projects/x-ees250129/cson-forge-data/source-data/UNIFIED_BGC/BGCdataset.nc
▶️ [1/8] Writing ROMS grid...
▶️ [2/8] Generating initial conditions...
/home/x-mlong/codes/cson-forge/cson_forge/_core.py:1370: UserWarning: Failed to compare grid datasets: [Errno 2] No such file or directory: '/Users/mclong/cson-forge-data/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_grid.nc'
if not self._file_blueprint_data_match(partition_files=partition_files) or clobber:
/home/x-mlong/.local/share/mamba/envs/cson-forge-v0/lib/python3.13/site-packages/dask/array/reductions.py:292: RuntimeWarning: All-NaN slice encountered
return np.nanmin(x_chunk, axis=axis, keepdims=keepdims)
[########################################] | 100% Completed | 80.89 s
▶️ [3/8] Generating surface forcing...
[########################################] | 100% Completed | 5.64 sms
▶️ [4/8] Generating surface forcing...
[########################################] | 100% Completed | 204.69 ms
▶️ [5/8] Generating boundary forcing...
[WARNING] The northern boundary is divided by land. It would be safer (but slower and more memory-intensive) to use `apply_2d_horizontal_fill = True`.
[########################################] | 100% Completed | 1.26 sms
▶️ [6/8] Generating boundary forcing...
[########################################] | 100% Completed | 30.77 s
▶️ [7/8] Generating tidal forcing...
[########################################] | 100% Completed | 6.73 sms
▶️ [8/8] Generating river forcing...
[WARNING] No records found at or after the end_time: 2024-01-02 00:00:00.
✅ All input files generated.
[INFO] 🛠️ Configuring ROMSSimulation
[INFO] 🔧 Setting up ROMSExternalCodeBase...
[INFO] 🔧 Setting up MARBLExternalCodeBase...
[INFO] 📦 Fetching compile-time code...
[INFO] 📦 Fetching runtime code...
[INFO] 📦 Fetching input datasets...
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_grid.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_initial_conditions.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_tidal.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_river.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_boundary-physics_202401.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_boundary-bgc_clim.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_surface-physics_202401.nc into (16,20)
[INFO] Partitioning /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/input_datasets/cson_roms-marbl_v0.1_ccs-12km_surface-bgc_clim.nc into (16,20)
CstarSpecBuilder(description='California Current System', model_name='cson_roms-marbl_v0.1', grid_name='ccs-12km', grid_kwargs={'nx': 224, 'ny': 440, 'size_x': 2688, 'size_y': 5280, 'center_lon': -134.5, 'center_lat': 39.6, 'rot': 33.3, 'N': 60, 'hc': 250.0, 'theta_s': 6.0, 'theta_b': 6.0, 'verbose': True, 'hmin': 5.0}, open_boundaries=OpenBoundaries(north=True, south=True, east=True, west=True), partitioning=PartitioningParameterSet(documentation='', locked=False, hash=None, n_procs_x=16, n_procs_y=20), start_date=datetime.datetime(2024, 1, 1, 0, 0), end_date=datetime.datetime(2024, 1, 2, 0, 0), cdr_forcing=None, blueprint=RomsMarblBlueprint(name='cson_roms-marbl_v0.1_ccs-12km', description='California Current System', application='roms_marbl', state='notset', valid_start_date='2024-01-01T00:00:00', valid_end_date='2024-01-02T00:00:00', code=ROMSCompositeCodeRepository(roms={'documentation': '', 'locked': False, 'location': 'https://github.com/CWorthy-ocean/ucla-roms.git', 'commit': '84f4ee7886e9ee4c33b3248b35c955551f3b9c06', 'branch': '', 'filter': None}, run_time=CodeRepository(documentation='', locked=False, location='/home/x-mlong/codes/cson-forge/cson_forge/builds/cson_roms-marbl_v0.1_ccs-12km/run-time', commit='', branch='na', filter={'files': ['marbl_diagnostic_output_list', 'marbl_in', 'marbl_tracer_output_list', 'roms.in']}), compile_time=CodeRepository(documentation='', locked=False, location='/home/x-mlong/codes/cson-forge/cson_forge/builds/cson_roms-marbl_v0.1_ccs-12km/compile-time', commit='', branch='na', filter={'files': ['Makefile', 'bgc.opt', 'blk_frc.opt', 'cdr_frc.opt', 'cppdefs.opt', 'diagnostics.opt', 'ocean_vars.opt', 'param.opt', 'river_frc.opt', 'sponge_tune.opt', 'surf_flux.opt', 'tides.opt', 'tracers.opt']}), marbl={'documentation': '', 'locked': False, 'location': 'https://github.com/marbl-ecosys/MARBL.git', 'commit': 'marbl0.45.0', 'branch': '', 'filter': None}), initial_conditions={'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_initial_conditions.nc', 'partitioned': False}]}, grid={'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_grid.nc', 'partitioned': False}]}, forcing={'boundary': {'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_boundary-physics_202401.nc', 'partitioned': False}, {'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_boundary-bgc_clim.nc', 'partitioned': False}]}, 'surface': {'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_surface-physics_202401.nc', 'partitioned': False}, {'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_surface-bgc_clim.nc', 'partitioned': False}]}, 'tidal': {'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_tidal.nc', 'partitioned': False}]}, 'river': {'documentation': '', 'locked': False, 'data': [{'location': '/anvil/projects/x-ees250129/cson-forge-data/x-mlong/input-data/cson_roms-marbl_v0.1_ccs-12km/cson_roms-marbl_v0.1_ccs-12km_river.nc', 'partitioned': False}]}, 'corrections': None}, partitioning={'documentation': '', 'locked': False, 'hash': None, 'n_procs_x': 16, 'n_procs_y': 20}, model_params={'time_step': 32}, runtime_params={'start_date': datetime.datetime(2024, 1, 1, 0, 0), 'end_date': datetime.datetime(2024, 1, 2, 0, 0), 'output_dir': PosixPath('/anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102')}, cdr_forcing=None), src_data=SourceData(datasets=['ERA5', 'GLORYS_REGIONAL', 'TPXO', 'UNIFIED_BGC'], clobber=False, grid=Grid(nx=224, ny=440, size_x=2688, size_y=5280, center_lon=-134.5, center_lat=39.6, rot=33.3, N=60, theta_s=6.0, theta_b=6.0, hc=250.0, topography_source={'name': 'ETOPO5'}, hmin=5.0, verbose=True, straddle=False), grid_name='ccs-12km', start_time=datetime.datetime(2024, 1, 1, 0, 0), end_time=datetime.datetime(2024, 1, 2, 0, 0)), grid=Grid(nx=224, ny=440, size_x=2688, size_y=5280, center_lon=-134.5, center_lat=39.6, rot=33.3, N=60, theta_s=6.0, theta_b=6.0, hc=250.0, topography_source={'name': 'ETOPO5'}, hmin=5.0, verbose=True, straddle=False))Run Simulation¶
Execute the model simulation. The run() method handles the execution and returns an execution handler for monitoring the run.
exec_handler = builder.run()
print(exec_handler)[INFO] Running srun -n 320 /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/compile_time_code/roms /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/input/runtime_code/cson_roms-marbl_v0.1_ccs-12km.in
[INFO] Submitting job: sbatch /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/work/cson_roms-marbl_v0-1_ccs-12km_20240101-20240102.sh
<cstar.execution.scheduler_job.SlurmJob object at 0x1506d270e900>
dot_count = 0
while not handler.ExecutionStatus.is_terminal(exec_handler.status):
print("...", end="", flush=True)
dot_count += 3
if dot_count >= 100:
print() # New line after ~100 dots
dot_count = 0
time.sleep(30)
if exec_handler.status == handler.ExecutionStatus.COMPLETED:
builder.post_run()
else:
raise Exception("Model run failed").....................................................................[INFO] Joining netCDF files output_bgc_dia.20240101000000.*.nc...
[INFO] Joining netCDF files output_bgc.20240101000000.*.nc...
[INFO] Joining netCDF files output_rst.20240101120000.*.nc...
[INFO] Joining netCDF files output_rst.20240102000000.*.nc...
[INFO] Joining netCDF files output_his.20240101000000.*.nc...
[INFO] done spatially joining /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/output/output_his.20240101000000.nc
[INFO] done spatially joining /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/output/output_bgc.20240101000000.nc
[INFO] done spatially joining /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/output/output_rst.20240101120000.nc
[INFO] done spatially joining /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/output/output_rst.20240102000000.nc
[INFO] done spatially joining /anvil/scratch/x-mlong/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_20240101-20240102/output/output_bgc_dia.20240101000000.nc
# Find the latest cstar log file in the run_path directory
log_files = sorted(glob(str(exec_handler.output_file)), reverse=True)
if log_files:
latest_log = Path(log_files[0])
print(f"Latest log file: {latest_log}")
print(f"Modified: {latest_log.stat().st_mtime}")
latest_log
else:
print("No cstar log files found")No cstar log files found
Visualize Model Output¶
After the model run completes, you can load and visualize the output data. The code below:
Finds output files: Uses
globto locate all BGC (biogeochemical) output files in theJOINED_OUTPUTdirectoryOpens the dataset: Uses
xarray.open_mfdataset()to open multiple NetCDF files as a single datasetApplies land mask: Masks out land points using the grid’s
mask_rhovariablePlots a variable: Creates a plot of dissolved inorganic carbon (DIC) at the first time step and bottom vertical level (
s_rho=-1)
The JOINED_OUTPUT directory contains the spatially-joined output files created by post_run(), which combine partitioned output files from parallel runs into single file
files = glob(str(builder.run_output_dir / "joined_output" / ("output_bgc.*")))
ds = xr.open_mfdataset(files)
ds = ds.where(builder.grid.ds.mask_rho)
ds.DIC.isel(time=0, s_rho=-1).plot()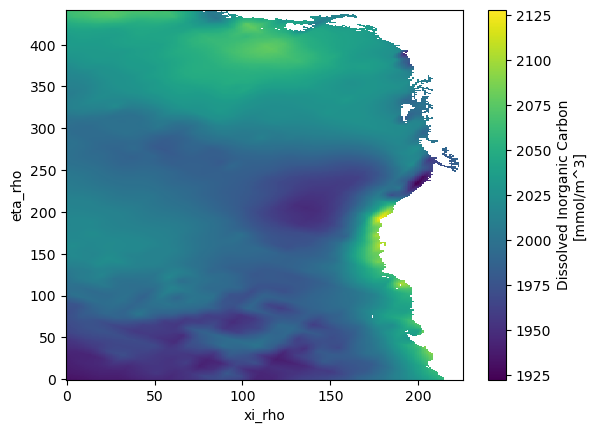
Save Executed Notebook¶
Save a timestamped copy of this notebook to executed/forge/{os}/ for reproducibility and record-keeping. The copy is organized by operating system (macOS or Ubuntu/Linux) to track execution history across different platforms.
The saved notebook includes all executed cells and outputs, providing a complete record of the simulation workflow for future reference.
# Save the notebook copy
cson_forge.save_notebook_copy(notebook_name="CStarSpecEngine-build-one.ipynb")Notebook copy saved to: executed/forge/RCAC_anvil/CStarSpecEngine-build-one_RCAC_anvil.ipynb
PosixPath('executed/forge/RCAC_anvil/CStarSpecEngine-build-one_RCAC_anvil.ipynb')