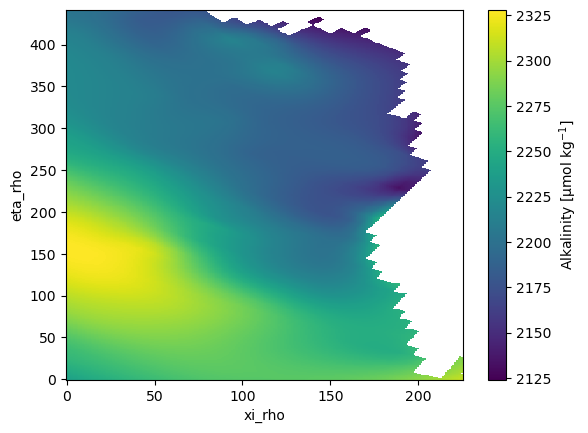
Compare simulated DIC and Alkalinity fields to observationally-based data
This notebook is intended to evaluate model skill by comparing simulated Dissolved Inorganic Carbon (DIC) and Total Alkalinity (ALK) fields against GLODAP observational products. The analysis focuses on spatial patterns and summary metrics that highlight biases relative to the GLODAP climatology.
%load_ext autoreload
%autoreload 2
from glob import glob
from pathlib import Path
import numpy as np
import xarray as xr
import xesmf
import cson_forge
from cson_forge.diagnostics import glodap
domain_name = "CCS-12km"
model_name = "cson_roms-marbl_v0.1"
grid_name = "ccs-12km"
dask_cluster_kwargs = {
"account": "m4632",
"queue_name": "premium",
"n_nodes": 1,
"n_tasks_per_node": 128,
"wallclock": "06:00:00",
"scheduler_file": None,
}
# Parameters
domain_name = "CCS-12km"
grid_name = "ccs-12km"
model_name = "cson_roms-marbl_v0.1"
dask_cluster_kwargs = {
"account": "m4632",
"queue_name": "premium",
"n_nodes": 1,
"n_tasks_per_node": 128,
"wallclock": "06:00:00",
"scheduler_file": None,
}
df = cson_forge.catalog.blueprint.load(stage="run")
df = df.loc[(df.grid_name == grid_name) & (df.model_name == model_name)]
if df.empty:
raise ValueError(
f"No blueprint found: model_name='{model_name}'; grid_name='{grid_name}' "
f"at stage='run'"
)
dfLoading...
blueprint_path = df.blueprint_path.iloc[0]
casename = blueprint_path.stem.replace("B_", "") # get less janky
simulation_dir = Path(cson_forge.config.paths.scratch / casename)
print(f"Blueprint path: {blueprint_path}")
print(f"Simulation directory: {simulation_dir}")
# TODO: make this less janky
pattern = "output_bgc_dia.*"
files = glob(str(simulation_dir / "output" / "joined_output" / pattern))
print(f"Found {len(files)} files matching pattern '{pattern}'")
if files:
print(f"First file: {Path(files[0]).name}")
if len(files) > 1:
print(f"Last file: {Path(files[-1]).name}")
print(f"All files: {[Path(f).name for f in sorted(files)]}")
else:
print("No files found!")Blueprint path: /Users/mclong/codes/cson-forge/cson_forge/blueprints/cson_roms-marbl_v0.1_ccs-12km/B_cson_roms-marbl_v0.1_ccs-12km_run_20240101-20240102.yml
Simulation directory: /Users/mclong/cson-forge-data/cson-forge-run/cson_roms-marbl_v0.1_ccs-12km_run_20240101-20240102
Found 1 files matching pattern 'output_bgc_dia.*'
First file: output_bgc_dia.20120101000000.0.nc
domain_name = f"{model_name}_{grid_name}"
grid_yaml = Path(df.grid_yaml_path.iloc[0])
print("Reading grid file:\n", grid_yaml)
grid_obj = cson_forge.parsers.load_roms_tools_object(grid_yaml)
grid = grid_obj.ds
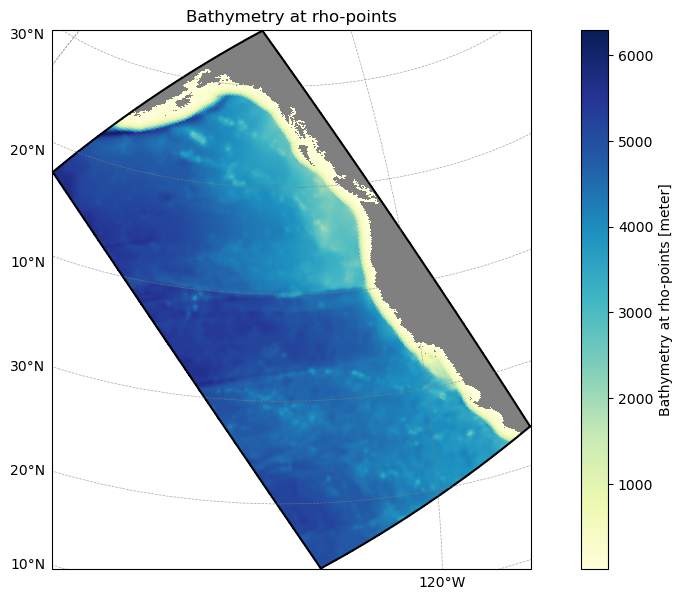
grid_obj.plot();Reading grid file:
/Users/mclong/codes/cson-forge/cson_forge/blueprints/cson_roms-marbl_v0.1_ccs-12km/_grid.yml
ds = glodap.open_glodap(product='GLODAPv2.2016b_MappedClimatologies')
# subset for surface
ds = ds.isel(depth=0, drop=True)
# Regrid GLODAP data to model grid using xesmf (grid loaded from cell above)
# Create source grid (GLODAP) - 2D lon/lat for xesmf
lon_2d, lat_2d = np.meshgrid(ds.lon.values, ds.lat.values)
ds_source = xr.Dataset(
{"lon": (("lat", "lon"), lon_2d), "lat": (("lat", "lon"), lat_2d)}
)
# Create destination grid (model grid)
ds_target = xr.Dataset(
{
"lon": (("eta_rho", "xi_rho"), grid.lon_rho.values),
"lat": (("eta_rho", "xi_rho"), grid.lat_rho.values),
}
)
# Build regridder (bilinear, periodic for global data)
regridder = xesmf.Regridder(
ds_source, ds_target, method="bilinear", periodic=True
)
# Variables to regrid (exclude coords and derived)
skip_vars = ["lon", "lat", "area", "depth_bnds", "dz"]
vars_to_regrid = [v for v in ds.data_vars if v not in skip_vars]
ds_to_regrid = ds[vars_to_regrid]
# Regrid
ds_regridded = regridder(ds_to_regrid)
# Add model grid coordinates and copy attributes
ds_regridded = ds_regridded.assign_coords(
lon_rho=(("eta_rho", "xi_rho"), grid.lon_rho.values),
lat_rho=(("eta_rho", "xi_rho"), grid.lat_rho.values),
)
for v in ds_regridded.data_vars:
if v in ds.data_vars:
ds_regridded[v].attrs.update(ds[v].attrs)
ds_regridded.attrs.update(ds.attrs)
ds = ds_regridded
print("GLODAP data regridded to model grid using xesmf (bilinear, periodic)")
ds/Users/mclong/.local/share/mamba/envs/cson-forge-v0/lib/python3.13/site-packages/xesmf/backend.py:42: UserWarning: Input array is not F_CONTIGUOUS. Will affect performance.
warnings.warn('Input array is not F_CONTIGUOUS. ' 'Will affect performance.')
GLODAP data regridded to model grid using xesmf (bilinear, periodic)
Loading...
ds.DIC.plot();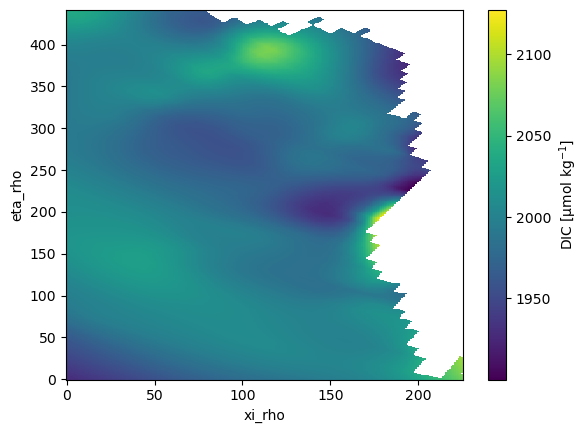
ds.ALK.plot();